Journal Club: Why are there so many bird species in the tropics?

SUMMARY: What can we learn about evolution, geography and biodiversity by studying continental patterns of speciation?
![]() Since before the time of dinosaurs, species diversity is related to latitude. Basically, species richness increases as distance from the equator decreases. As any sweaty bird watcher dragging a heavy field guide through the tropics will tell you, this is especially obvious for New World birds where 3,370 species breed in the Neotropics whilst only 732 species breed in the Nearctic. Even Charles Darwin commented on this latitudinal diversity gradient.
Since before the time of dinosaurs, species diversity is related to latitude. Basically, species richness increases as distance from the equator decreases. As any sweaty bird watcher dragging a heavy field guide through the tropics will tell you, this is especially obvious for New World birds where 3,370 species breed in the Neotropics whilst only 732 species breed in the Nearctic. Even Charles Darwin commented on this latitudinal diversity gradient.
Hypotheses abound — hypotheses with hairsplitting and convoluted minutiæ (sometimes known as “arm waving”) that usually give me a migraine — yet the biogeographical reasons underlying this curious phenomenon remain elusive. But that said, the most widely accepted explanation for this tremendous latitudinal diversity gradient in bird species is the refuge hypothesis. This hypothesis proposes that the climate and habitat changes that occurred due to the glacial cycles that started during the late Pliocene and continued through Pleistocene caused bird populations to become isolated into small refugia, and this population fragmentation triggered increased speciation rates during that particular time period — a rapid rate that still continues to this day. According to this hypothesis, it was this rapid diversification that created the high Neotropical bird diversity seen today.
Another explanation for this avian latitudinal diversity gradient is the climate stability hypothesis. This hypothesis suggests that the stable environment found in the tropics provided birds with the opportunity to specialize on predictable resources, thus allowing them to occupy narrower niches and this in turn, facilitated speciation.
Both hypotheses sound reasonable, so how can one decide which of the two is more likely? One way to figure this out is to test each hypothesis’s predictions. Because the refuge hypothesis suggests there is greater evolutionary potential in the tropics, one would expect that tropical birds consistently speciate rapidly, so tropical species should be younger than those found in temperate regions. In contrast, the climate stability hypothesis predicts that species found in high biodiversity (tropical) regions speciated early, filling up all available niches, and therefore should be older than birds found in low biodiversity (temperate) regions, which had the opportunity to speciate only recently, after the glaciers disappeared.
To test these hypotheses, it is necessary to determine two things: how closely related are neighboring congeners (birds in the same genus that are geographically close), and how closely related are sister species (a species pair derived from a common ancestor shared by no other species). For many decades, scientists lacked the technology to accurately determine these parameters on a large scale, but the advent of DNA barcoding has changed this. Originally, DNA barcoding was used purely for identification purposes. This technique examines a small genetic region of a widely-shared gene to detect small changes and uses those to identify species. The most commonly used region, or “marker”, for animals is the cytochrome c oxidase I (COI) gene, and thousands of these sequences are publicly available in scientific databases such as GenBank.
But DNA barcoding appears to be more versatile than originally thought. For example, even though it is controversial, comparisons and analyses of large collections of DNA barcoding data (DNA barcode libraries) may provide useful insights into differences in speciation rates and diversification patterns at the continental scale — just the sort of data needed to test these two hypotheses.
An international team of scientists assembled their own DNA barcode library for approximately 50% of the Argentine avifauna, approximately 50% of Palearctic bird species and a nearly complete dataset for North American birds. They then analyzed these data and compared genetic distances between neighboring congeners (figure 1):
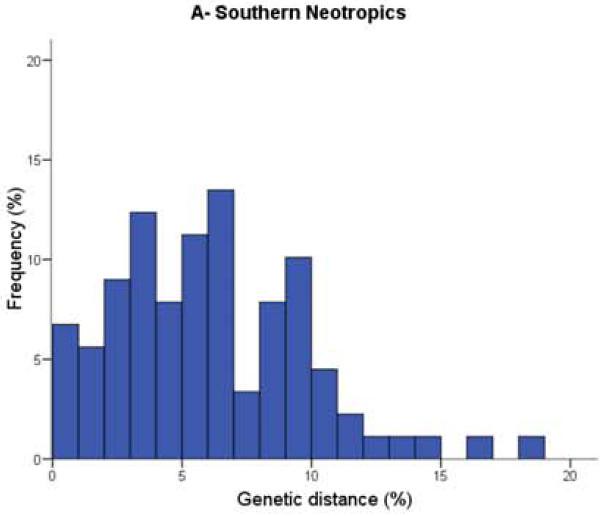
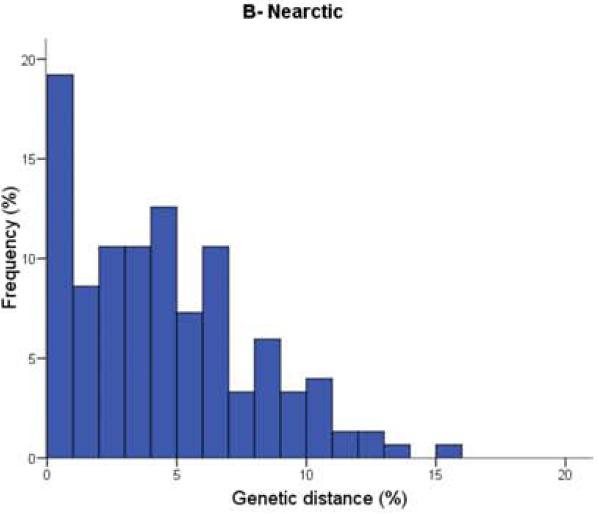
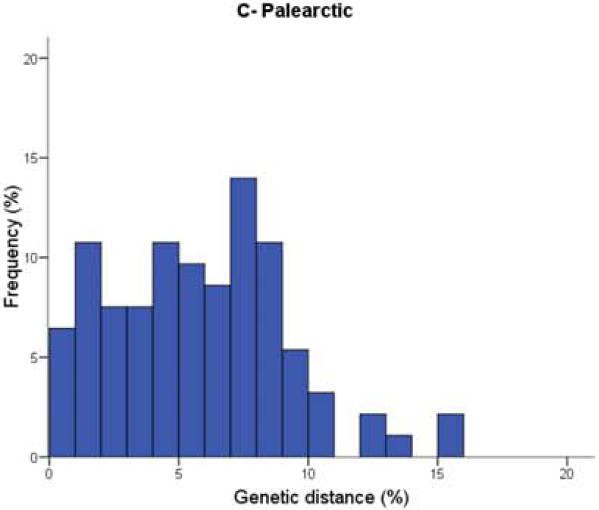
Figure 1. Distribution of nearest congeneric neighbour distances in COI for bird species in three biogeographic regions.
DOI: 10.1371/journal.pone.0020744
They found the distribution of genetic distances (x-axis) between nearest congeneric neighbours was significantly different between birds in the southern Neotropics (Figure 1A: embiggen) and those in the Nearctic (Figure 1B: embiggen). As you can see, genetic distances were much smaller for Nearctic birds; species pairs with very low (less than 1%) divergence were three times higher in North America (18.5%) than in Argentina (only 6.7%). Comparing these data also showed that divergence values in the Palearctic (Figure 1C: embiggen) are closer to those of the Neotropics than to those in the Nearctic.
The team then conducted the same analysis and comparisons of divergence values for sister species (Figure 2):


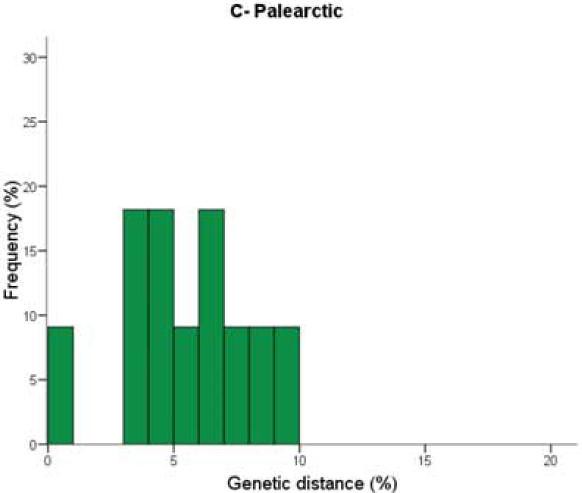
Figure 2. Distribution of sister species distances in COI for bird species in three biogeographic regions.
DOI: 10.1371/journal.pone.0020744
Once again, they found that the average genetic distances between sister species in southern Neotropics (Figure 2A: embiggen) were larger than those for Nearctic birds (Figure 2B: embiggen). (Note that the pattern for Palearctic birds [Figure 2C: embiggen] once again resembles that for Neotropical birds).
But what does this mean in years? Using the sister species dataset, the team compared divergence rates of COI to that of another mitochondrial gene, cytochrome b. Cytb is well-characterised and its rate of change is estimated to be 2.1% per million years across all species. Based on this comparison, the team estimated that the rate of change for COI is 1.4% per million years. Using this calibration as a molecular clock, they estimate that almost one-third of the Nearctic avian sister species pairs are quite young, appearing in just the last 700,000 years (mid to late Pleistocene). By contrast, considerably fewer of the southern Neotropic (15%) and Palearctic (10%) sister species pairs diverged so recently.
These data also indicate that Pleistocene glaciation cycles had a similarly mild effect upon Palearctic species as upon those in the Neotropics. Unfortunately, there were less data available for Palearctic species and fewer species pairs were analyzed, so this study only suggests a trend for those species examined.
The relative youth of North American temperate zone bird species indicates that the Pleistocene cycles of glaciation had a major effect on the Nearctic, but only a mild effect in the tropics. These data are consistent with the climate stability hypothesis, and contrary to the scientific consensus (although don’t forget that “consensus” doesn’t mean we all agree that the refuge hypothesis accurately explains this disparity in geographical biodiversity: getting scientists to agree 100% on anything is like herding cats, after all!).
These data also suggest that DNA barcoding is worth a closer look when trying to decipher the historic origins for the complex web of biodiversity patterns that we see on Earth today.
Source:
Lijtmaer, D., Kerr, K., Barreira, A., Hebert, P., & Tubaro, P. (2011). DNA Barcode Libraries Provide Insight into Continental Patterns of Avian Diversification. PLoS ONE, 6 (7) DOI: 10.1371/journal.pone.0020744
Background:
Mittelbach, G., Schemske, D., Cornell, H., Allen, A., Brown, J., Bush, M., Harrison, S., Hurlbert, A., Knowlton, N., Lessios, H., McCain, C., McCune, A., McDade, L., McPeek, M., Near, T., Price, T., Ricklefs, R., Roy, K., Sax, D., Schluter, D., Sobel, J., & Turelli, M. (2007). Evolution and the latitudinal diversity gradient: speciation, extinction and biogeography. Ecology Letters, 10 (4), 315-331 DOI: 10.1111/j.1461-0248.2007.01020.x
Jun Chen, Qi Li, Lingfeng Kong, & Hong Yu (2011). How DNA Barcodes Complement Taxonomy and Explore Species Diversity: The Case Study of a Poorly Understood Marine Fauna. PLoS ONE : 10.1371/journal.pone.0021326
Moritz, C., & Cicero, C. (2004). DNA Barcoding: Promise and Pitfalls. PLoS Biology, 2 (10) DOI: 10.1371/journal.pbio.0020354
Hebert, P., Stoeckle, M., Zemlak, T., & Francis, C. (2004). Identification of Birds through DNA Barcodes. PLoS Biology, 2 (10) DOI: 10.1371/journal.pbio.0020312
.. .. .. .. .. .. .. .. .. .. .. ..
twitter: @GrrlScientist
facebook: grrlscientist
email: grrlscientist@gmail.com
Top Photo: Red knot, Calidris canutus rufa, at Mispillion Harbor, Delaware (USA). This species winters in Argentina and makes an epic migration through North America to its arctic breeding grounds in Canada. NOTE: This image has been cropped from the original.
Image: Greg Breese/USFWS, 2 June 2008 (Public Domain) [velociraptorise].

Hi GrrlScientist, nice digest, I read it in the Guardian earlier https://www.guardian.co.uk/science/punctuated-equilibrium/2011/aug/18/2! Two points/corrections, 1) the refuge hypothesis has been ‘widely dicredited’ already as a driver of richness in the Neotropics (and was never a ‘consensus’) – where its all about fluvial barriers and temporal stability… and 2) C. c. rufa dosn’t breed in the Palearctic (it breeds in the Canadian Arctic).
ATB
Alex